The common theme of our research is to apply systems engineering strategies and control engineering principles and techniques to understand, predict, and control complex dynamic processes. Our research focuses on two groups of complex dynamic systems: microbiome and large-scale industrial processes. By developing new modeling strategies, wet lab protocols, and bioreactors, we tackle both fundamental and applied problems that we encounter in various industries relevant to climate change.
Harnessing Microbiome for Sustainability
In nature, almost all microorganisms exist in complex microbial communities. It is the interactions among different members that stabilize the structure and functionalities of the communities despite different environmental stresses. To harness synergistic interactions in the microbial community while keeping the microbiome tractable, we start with binary synthetic microbiomes.
To elucidate the emergent inter-species interactions that are difficult to measure directly, we developed an integrated computational-experimental strategy. The computational approach integrates unstructured kinetic modeling with structured genome-scale modeling; the experimental approach complements bioreactor growth data with meta-transcriptomic data. An iterative procedure between computation and experimentation enables us to gain a better understanding of how different environmental and genetic factors determine the structure and functionality of the synthetic microbiome. One focus of this area is the methanotroph-photoautotroph coculture for biogas conversion.
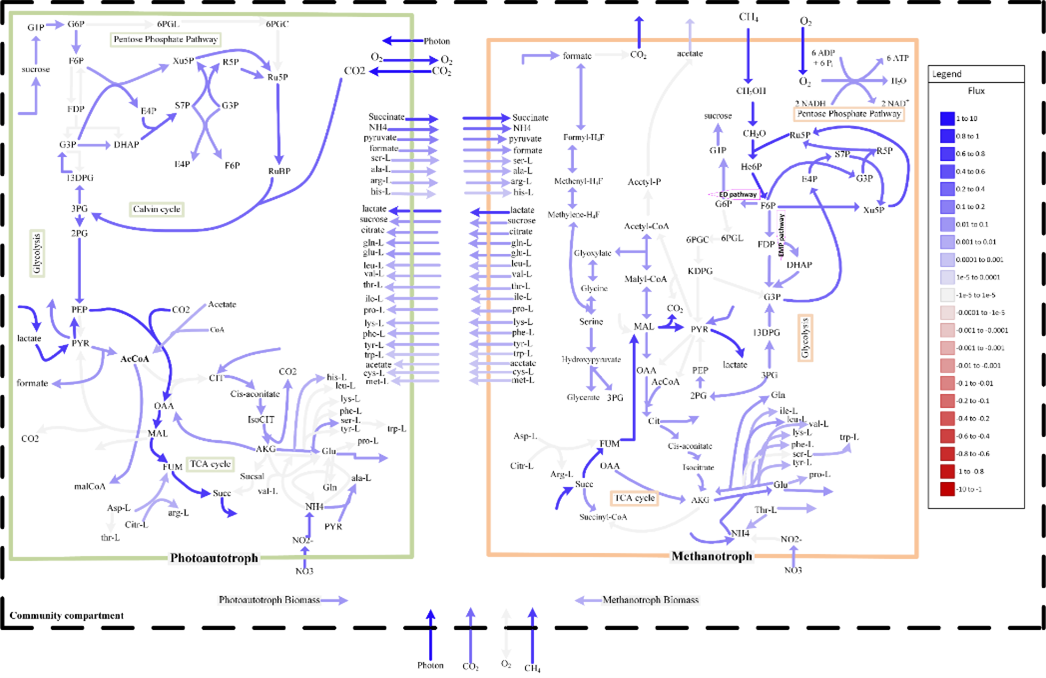
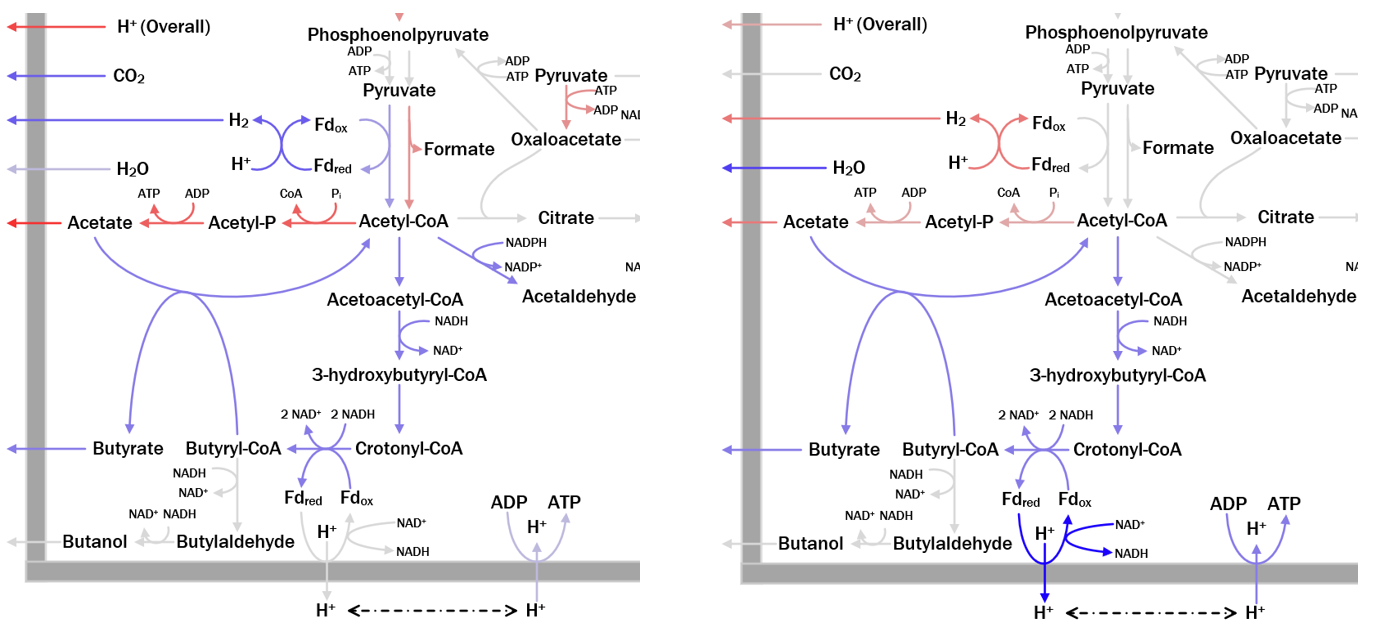
A System Identification Framework for Genome-Scale Metabolic Modeling and Analysis
A high-quality genome-scale metabolic model (GEM) can help answer many questions about the organism. However, obtaining a high-quality GEM is highly time-consuming, even with the introduction of several genome-scale reconstruction tools that offer automated draft network generation and gap filling. To address this challenge, we have developed a system-identification based framework for GEM development and analysis. The framework deploys a knowledge-matching based computation framework for GEM analysis and refinement and has been successfully applied to develop and refine GEMs for different microbes: methanotroph, cyanobacteria, microalgae, and Clostridia strains.
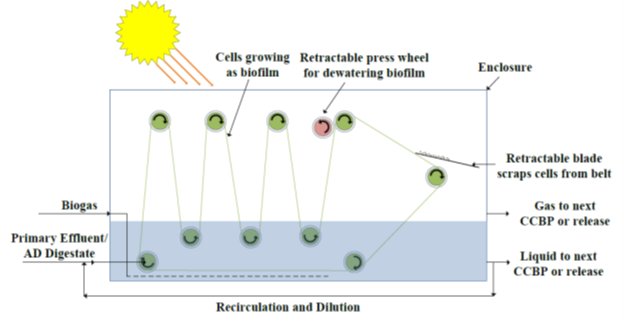
Biofilm-Based Cultivation for Gas Substrate Conversion
Biological conversion of gas substrates (especially the ones with small solubility such as methane, carbon monoxide, and oxygen) is inherently challenging due to mass transfer resistance. These processes usually suffer from low volumetric or areal productivity and require a large footprint to achieve the desired throughput. In addition, biological conversion involving photosynthesis suffers the same fate due to limited light penetration in the liquid phase. To address these challenges, we explore biofilm-based cultivation as a new platform for biotechnology by designing a novel biofilm-based bioreactor or photobioreactor with application in methane capture from air and biogas valorization.
Smart Manufacturing with Artificial Intelligence
Digitalization and automation are the game changers for Industrial 4.0. Due to the advances in sensor, control and information technologies, industries of all sizes and all types are sitting on mountains of data. It is crucial to extract meaningful information from large and complex data sets in order to assess process performance and improve process efficiency and product quality. By integrating human learning and machine learning, we are developing new approaches to extract key features from complex big data to accurately characterize the process behavior. The feature-based modeling enables effective monitoring, control, and predictive maintenance of different industrial processes. The applications in this area cover the chemical industry, pulp and paper industry, and semiconductor industry.